Image Preprocessing & Contour Extraction
José Luis Otero-Ferrer & Amalia Manjabacas
2026-05-19
Source:vignettes/Obtaining_Contour.Rmd
Obtaining_Contour.RmdAbout this tutorial
This tutorial describes how to use the aforoR package to process images, extract otolith contours, and calculate shape descriptors including Elliptic Fourier Descriptors (EFDs), Wavelets, and morphometrics variables.
1. Image preparation and Standardizing
Otolith images should be placed with the sulcus acusticus facing upward, the rostrum to the right, and the dorsal margin at the top (Tuset et al., 2008; Lombarte and Tuset, 2015), according to its natural position.
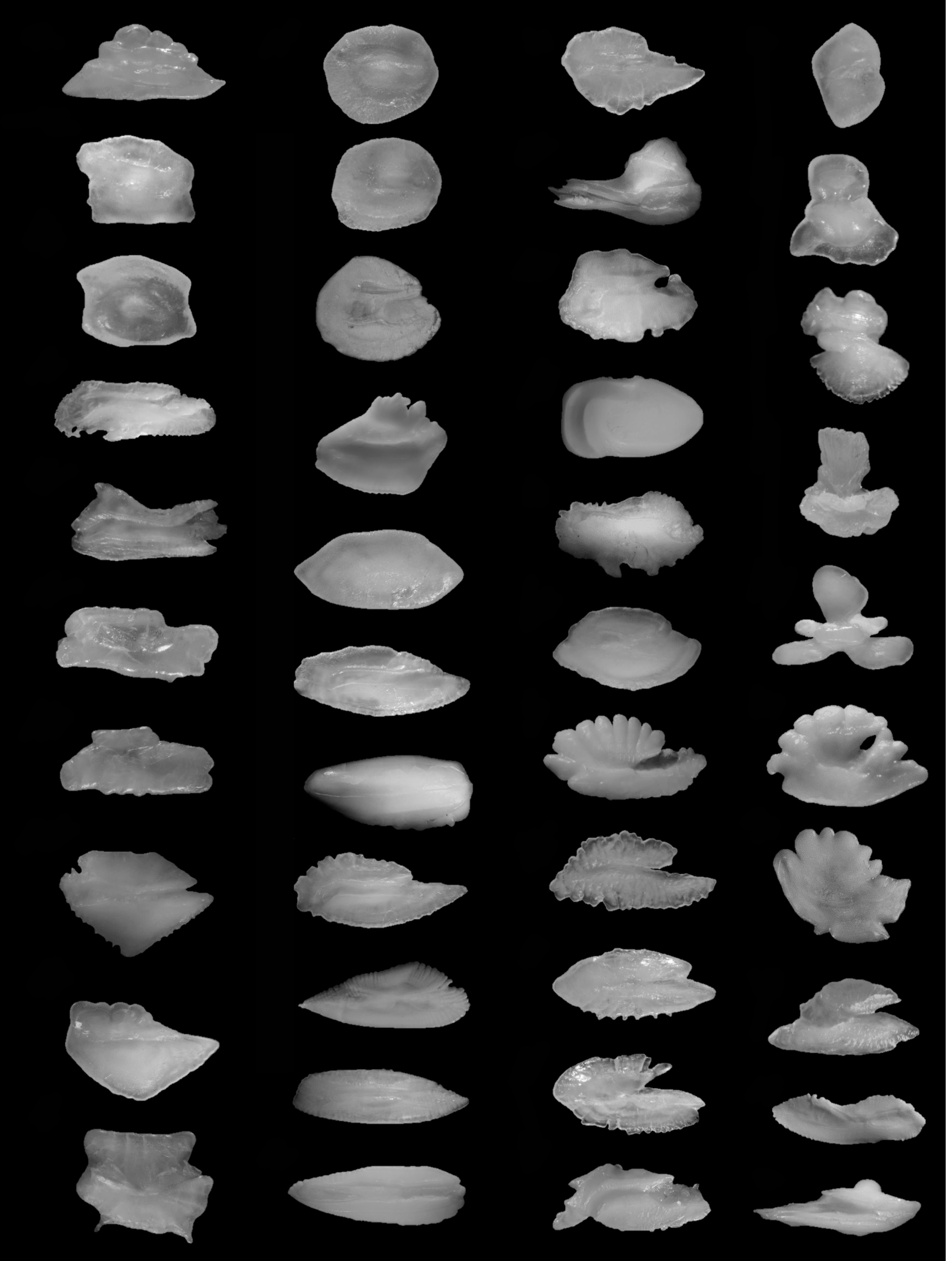

During image acquisition, modeling clay can be helpful for otoliths with pronounced convexity.
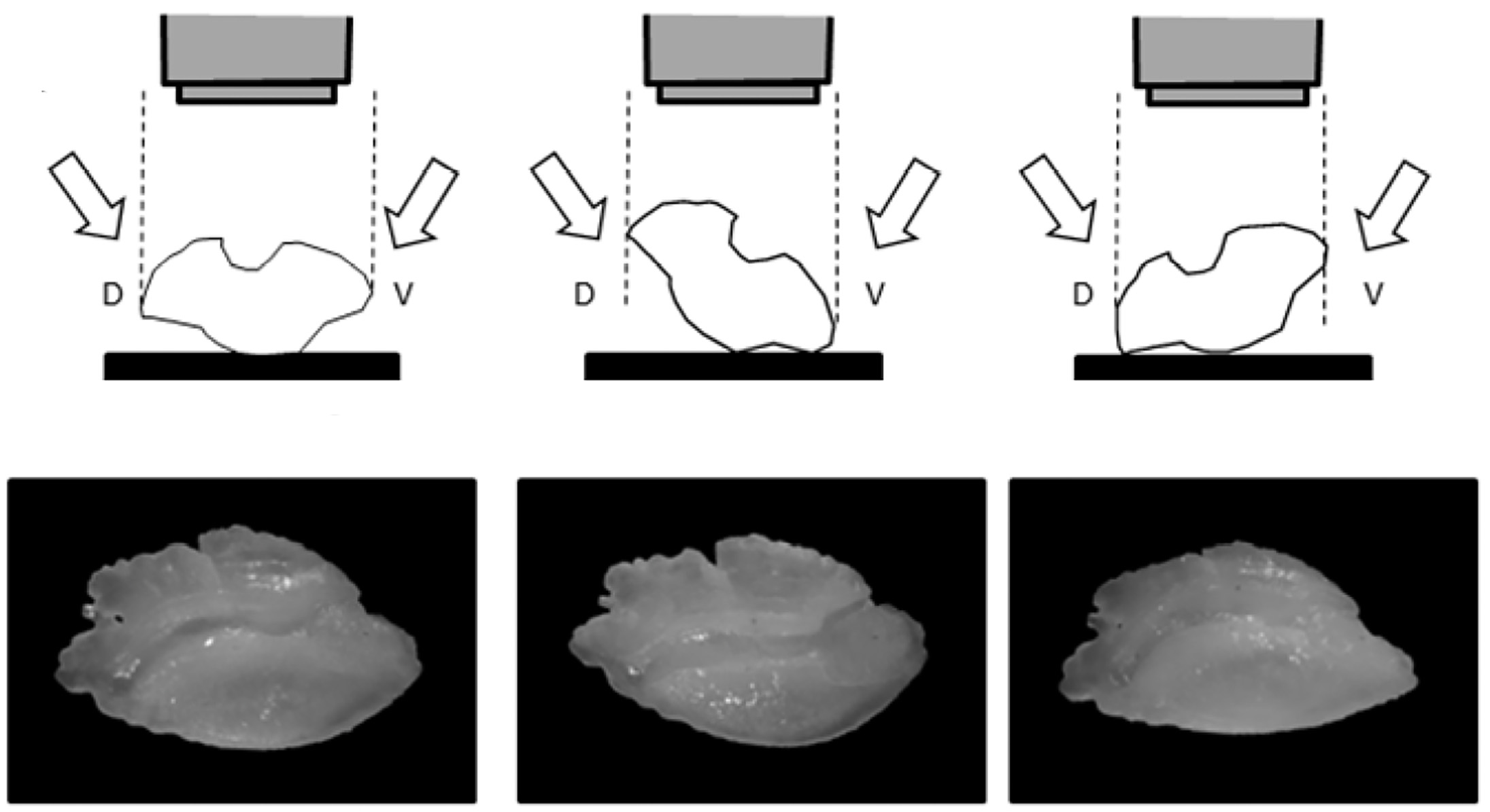
The package determine automatically the initial contour point, which represents the maximum distance from the otolith centroid to this point locating to the right. The rostrum is usually the reference point; however, in otoliths where the rostrum is not clearly defined, it is preferable to select another homologous point, as illustrated in the following example.
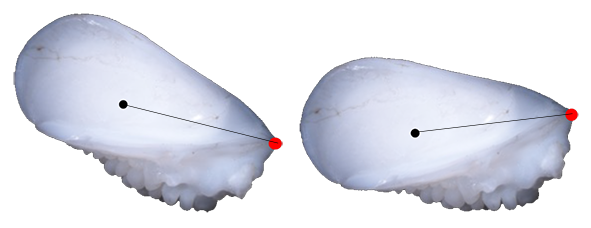
Automatic contour detection is highly sensitive to illumination conditions. To minimize variability and ensure reliable results, we recommend acquiring all images under consistent lighting and background conditions (preferably uniform black color).
2. Setting up images
The package works by processing all supported image files
(.jpg, .jpeg, .png,
.tif, .tiff) within a specified folder. While
we use sample images for this tutorial, full datasets of Aphanopus
carbo and A. intermedius (as used in other vignettes) can
be downloaded from Zenodo:
[!NOTE] Zenodo Dataset: Aphanopus Otolith Images
2.1 Folder Organization Best Practices
To ensure mathematically and biologically sound analyses, it is highly recommended to organize your image dataset structurally before running the package. Place images of different species, populations, or stocks into separate, dedicated subfolders.
For example:
Project_Images/
├── Aphanopus_carbo_Azores/
│ ├── otolith1.jpg
│ └── otolith2.jpg
└── Aphanopus_intermedius_Madeira/
├── otolith3.jpg
└── otolith4.jpgThis structure is particularly critical if you intend to use the
Generalized Procrustes Analysis (GPA) alignment
(procrustes = TRUE). GPA calculates a global “consensus
shape” for all specimens in the folder. If you mix highly divergent
species in the same folder, the algorithm will attempt to find a
non-biological average between them, distorting your morphometric
signals. By processing species/stocks homogeneously in their respective
folders, Procrustes operates optimally.
Furthermore, keeping datasets physically segregated simplifies data
management. Although aforoR will generate separate output
tables (.csv) for each folder, you can easily bind them
together in R downstream, automatically assigning species or stock
labels based on the directory names.
2.2 Setting up the working directory
First, let’s look at where the example image is stored:
library(aforoR)
# Locate the example image
image_path <- system.file("extdata", "otolith.jpg", package = "aforoR")
# Fallback if the package is not installed but we are running vignettes locally
if (image_path == "") {
image_path <- file.path("../inst/extdata", "otolith.jpg")
}
# Create a temporary directory for processing
work_dir <- file.path(tempdir(), "aforo_tutorial")
if (dir.exists(work_dir)) unlink(work_dir, recursive = TRUE)
dir.create(work_dir)
# Copy the example image to the working directory
file.copy(image_path, file.path(work_dir, "otolith.jpg"))
#> [1] TRUE
cat("Working directory:", work_dir, "\n")
#> Working directory: /tmp/RtmpfXBinh/aforo_tutorial
list.files(work_dir)
#> [1] "otolith.jpg"3. Processing images
The main function process_images handles the entire
workflow: 1. Preprocessing: Grayscale conversion,
filtering, and binarization. 2. Contour Extraction:
Identifies the otolith boundaires. 3. Feature
Calculation: Computes distances, wavelets, and EFDs. 4.
Morphometrics: Calculates linear measurements and
relative indices.
You can specify the scale of your images using the
pixels_per_mm argument to get measurements in
millimeters.
# Run the processing pipeline
# We set pixels_per_mm = 100 as an example scale (100 pixels = 1 mm)
process_images(
folder = work_dir,
threshold = NULL, # Automatic thresholding (Otsu)
wavelets = TRUE, # Calculate wavelets
ef = TRUE, # Calculate Elliptic Fourier Descriptors
pixels_per_mm = 100 # define scale
)
#>
#> Phase 1: Extracting contours...
#> | | | 0% | |========================================| 100%
#>
#> Phase 2: Calculating descriptors and saving results...
#> | | | 0% | |========================================| 100%4. Examining Results
The function creates two subdirectories: Polar (Polar
coordinates) and Cartesian (Perimeter coordinates).
4.1 Morphometric Measurements
A new file MorphometricsEN.csv is created containing
geometric measurements.
morpho_file <- file.path(work_dir, "Polar", "MorphometricsEN.csv")
if (file.exists(morpho_file)) {
morpho_data <- read.table(morpho_file, header = TRUE, sep = ";", dec = ".")
knitr::kable(morpho_data, caption = "Morphometric Indices")
}| Image | Area | Perimeter | Length | Width | Feret_Max | Feret_Min | PCA_Angle | Units |
|---|---|---|---|---|---|---|---|---|
| otolith.jpg | 9.7628 | 12.89881 | 5.025411 | 2.72705 | 5.031948 | 2.726763 | 3.082376 | mm |
The measurements include:
- Area: Area of the otolith ().
- Perimeter: Perimeter length ().
- Length / Width: Major and minor axis dimensions of the PCA-aligned bounding box ().
- Feret_Max: Maximum Feret diameter (longest distance between any two boundary points; recommended robust measure of length) ().
- Feret_Min: Minimum Feret diameter (minimum distance between parallel tangent lines; recommended robust measure of width) ().
- PCA_Angle: Orientation angle of the major axis of inertia in degrees (normalized to [0, 180)).
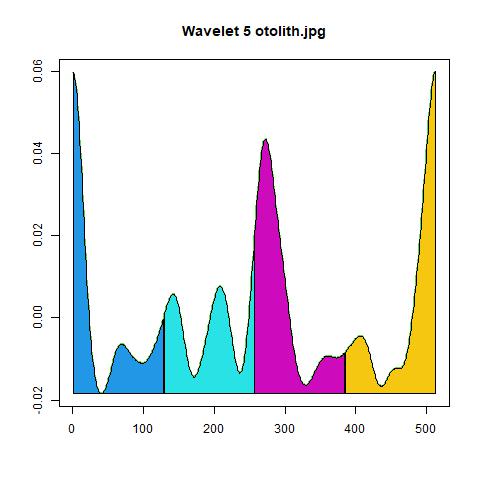
4.2 Wavelet Coefficients
The wavelet analysis results are saved in multiple CSV files corresponding to different scales.
wavelet_file <- file.path(work_dir, "Polar", "Wavelet_5EN.csv")
if (file.exists(wavelet_file)) {
wave5 <- read.table(wavelet_file, header = FALSE, sep = ";", dec = ".")
# Display first few columns
knitr::kable(wave5[, 1:10], caption = "Wavelet Scale 5 (First 10 columns)")
}| V1 | V2 | V3 | V4 | V5 | V6 | V7 | V8 | V9 | V10 |
|---|---|---|---|---|---|---|---|---|---|
| otolith.jpg | 0.0597147 | 0.0592524 | 0.0584312 | 0.0572526 | 0.055724 | 0.0538587 | 0.0516752 | 0.0491968 | 0.0464515 |
4.3 Visualizations
The package automatically generates diagnostic images to verify the contour extraction and analysis zones.